Alibaba.com offers 72 medical transcription products. About 40% of these are other business services, 2% are medical science. A wide variety of medical transcription options are available to you, There are 9 medical transcription suppliers, mainly located in Asia. PDF| Currently, prediction of transcription factor binding sites is widely done using matrices collected from literature. This leads to several problems. We cannot actively control the. PROMO is a virtual laboratory for the identification of putative transcription factor binding sites (TFBS) in DNA sequences from a species or groups of species of interest.
Transcription is the first step of DNA based gene expression, in which a particular segment of DNA is copied into RNA (especially mRNA) by the enzymeRNA polymerase.Both DNA and RNA are nucleic acids, which use base pairs of nucleotides as a complementary language. During transcription, a DNA sequence is read by an RNA polymerase, which produces a complementary, antiparallel RNA strand called a primary transcript.
Transcription proceeds in the following general steps:
- RNA polymerase, together with one or more general transcription factors, binds to promoter DNA.
- RNA polymerase creates a transcription bubble, which separates the two strands of the DNA helix. This is done by breaking the hydrogen bonds between complementary DNA nucleotides.
- RNA polymerase adds RNA nucleotides (which are complementary to the nucleotides of one DNA strand).
- RNA sugar-phosphate backbone forms with assistance from RNA polymerase to form an RNA strand.
- Hydrogen bonds of the RNA–DNA helix break, freeing the newly synthesized RNA strand.
- If the cell has a nucleus, the RNA may be further processed. This may include polyadenylation, capping, and splicing.
- The RNA may remain in the nucleus or exit to the cytoplasm through the nuclear pore complex.
The stretch of DNA transcribed into an RNA molecule is called a transcription unit and encodes at least one gene. If the gene encodes a protein, the transcription produces messenger RNA (mRNA); the mRNA, in turn, serves as a template for the protein's synthesis through translation. Alternatively, the transcribed gene may encode for non-coding RNA such as microRNA, ribosomal RNA (rRNA), transfer RNA (tRNA), or enzymatic RNA molecules called ribozymes.[1] Overall, RNA helps synthesize, regulate, and process proteins; it therefore plays a fundamental role in performing functions within a cell.
In virology, the term may also be used when referring to mRNA synthesis from an RNA molecule (i.e., RNA replication). For instance, the genome of a negative-sense single-stranded RNA (ssRNA -) virus may be template for a positive-sense single-stranded RNA (ssRNA +)[clarification needed]. This is because the positive-sense strand contains the information needed to translate the viral proteins for viral replication afterwards. This process is catalyzed by a viral RNA replicase.[2][clarification needed]
- 2Major steps
Background[edit]
A DNA transcription unit encoding for a protein may contain both a coding sequence, which will be translated into the protein, and regulatory sequences, which direct and regulate the synthesis of that protein. The regulatory sequence before ('upstream' from) the coding sequence is called the five prime untranslated region (5'UTR); the sequence after ('downstream' from) the coding sequence is called the three prime untranslated region (3'UTR).[1]
As opposed to DNA replication, transcription results in an RNA complement that includes the nucleotide uracil (U) in all instances where thymine (T) would have occurred in a DNA complement.
Only one of the two DNA strands serve as a template for transcription. The antisense strand of DNA is read by RNA polymerase from the 3' end to the 5' end during transcription (3' → 5'). The complementary RNA is created in the opposite direction, in the 5' → 3' direction, matching the sequence of the sense strand with the exception of switching uracil for thymine. This directionality is because RNA polymerase can only add nucleotides to the 3' end of the growing mRNA chain. This use of only the 3' → 5' DNA strand eliminates the need for the Okazaki fragments that are seen in DNA replication.[1] This also removes the need for an RNA primer to initiate RNA synthesis, as is the case in DNA replication.
The non-template (sense) strand of DNA is called the coding strand, because its sequence is the same as the newly created RNA transcript (except for the substitution of uracil for thymine). This is the strand that is used by convention when presenting a DNA sequence.[3]
Transcription has some proofreading mechanisms, but they are fewer and less effective than the controls for copying DNA. As a result, transcription has a lower copying fidelity than DNA replication.[4]
Major steps[edit]

Transcription is divided into initiation, promoter escape, elongation, and termination.[5]
Initiation[edit]
Transcription begins with the binding of RNA polymerase, together with one or more general transcription factors, to a specific DNA sequence referred to as a 'promoter' to form an RNA polymerase-promoter 'closed complex'. In the 'closed complex' the promoter DNA is still fully double-stranded.[5]
RNA polymerase, assisted by one or more general transcription factors, then unwinds approximately 14 base pairs of DNA to form an RNA polymerase-promoter 'open complex'. In the 'open complex' the promoter DNA is partly unwound and single-stranded. The exposed, single-stranded DNA is referred to as the 'transcription bubble.'[5]
RNA polymerase, assisted by one or more general transcription factors, then selects a transcription start site in the transcription bubble, binds to an initiating NTP and an extending NTP (or a short RNA primer and an extending NTP) complementary to the transcription start site sequence, and catalyzes bond formation to yield an initial RNA product.[5]
In bacteria, RNA polymerase holoenzyme consists of five subunits: 2 α subunits, 1 β subunit, 1 β' subunit, and 1 ω subunit. In bacteria, there is one general RNA transcription factor known as a sigma factor. RNA polymerase core enzyme binds to the bacterial general transcription (sigma) factor to form RNA polymerase holoenzyme and then binds to a promoter.[5](RNA polymerase is called a holoenzyme when sigma subunit is attached to the core enzyme which is consist of 2 α subunits, 1 β subunit, 1 β' subunit only).
In archaea and eukaryotes, RNA polymerase contains subunits homologous to each of the five RNA polymerase subunits in bacteria and also contains additional subunits. In archaea and eukaryotes, the functions of the bacterial general transcription factor sigma are performed by multiple general transcription factors that work together.[5] In archaea, there are three general transcription factors: TBP, TFB, and TFE. In eukaryotes, in RNA polymerase II-dependent transcription, there are six general transcription factors: TFIIA, TFIIB (an ortholog of archaeal TFB), TFIID (a multisubunit factor in which the key subunit, TBP, is an ortholog of archaeal TBP), TFIIE (an ortholog of archaeal TFE), TFIIF, and TFIIH. In archaea and eukaryotes, the RNA polymerase-promoter closed complex is usually referred to as the 'preinitiation complex.'[6]
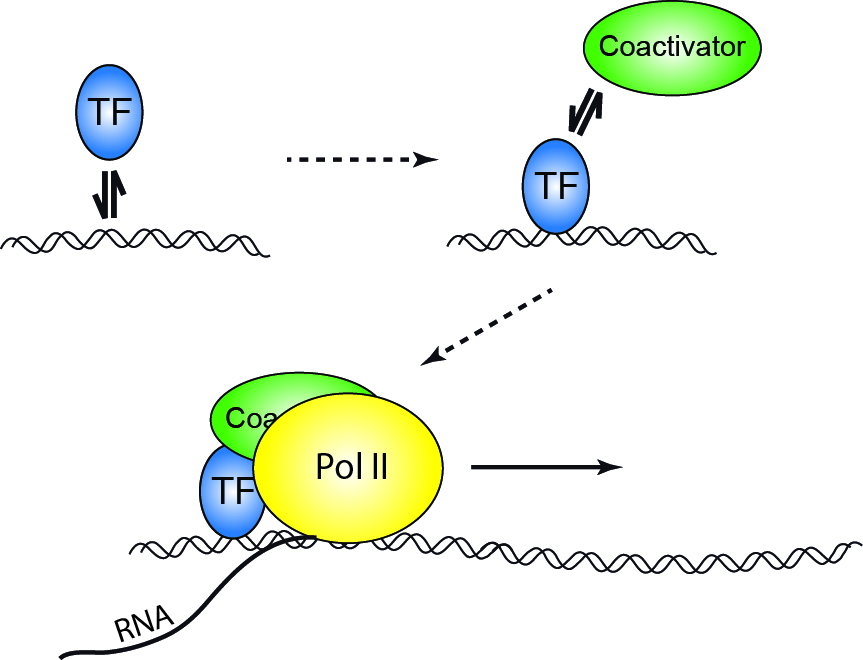
Transcription initiation is regulated by additional proteins, known as activators and repressors, and, in some cases, associated coactivators or corepressors, which modulate formation and function of the transcription initiation complex.[5]
Promoter escape[edit]
After the first bond is synthesized, the RNA polymerase must escape the promoter. During this time there is a tendency to release the RNA transcript and produce truncated transcripts. This is called abortive initiation, and is common for both eukaryotes and prokaryotes.[7] Abortive initiation continues to occur until an RNA product of a threshold length of approximately 10 nucleotides is synthesized, at which point promoter escape occurs and a transcription elongation complex is formed.
Mechanistically, promoter escape occurs through DNA scrunching, providing the energy needed to break interactions between RNA polymerase holoenzyme and the promoter.[8]
In bacteria, it was historically thought that the sigma factor is definitely released after promoter clearance occurs. This theory had been known as the obligate release model however later data showed that upon and following promoter clearance, the sigma factor is released according to a stochastic model known as the stochastic release model.[9]
In eukaryotes, at an RNA polymerase II-dependent promoter, upon promoter clearance, TFIIH phosphorylates serine 5 on the carboxy terminal domain of RNA polymerase II, leading to the recruitment of capping enzyme (CE).[10][11] The exact mechanism of how CE induces promoter clearance in eukaryotes is not yet known.
Elongation[edit]
One strand of the DNA, the template strand (or noncoding strand), is used as a template for RNA synthesis. As transcription proceeds, RNA polymerase traverses the template strand and uses base pairing complementarity with the DNA template to create an RNA copy (which elongates during the traversal). Although RNA polymerase traverses the template strand from 3' → 5', the coding (non-template) strand and newly formed RNA can also be used as reference points, so transcription can be described as occurring 5' → 3'. This produces an RNA molecule from 5' → 3', an exact copy of the coding strand (except that thymines are replaced with uracils, and the nucleotides are composed of a ribose (5-carbon) sugar where DNA has deoxyribose (one fewer oxygen atom) in its sugar-phosphate backbone).[citation needed]
mRNA transcription can involve multiple RNA polymerases on a single DNA template and multiple rounds of transcription (amplification of particular mRNA), so many mRNA molecules can be rapidly produced from a single copy of a gene.[citation needed] The characteristic elongation rates in prokaryotes and eukaryotes are about 10-100 nts/sec.[12] In eukaryotes, however, nucleosomes act as major barriers to transcribing polymerases during transcription elongation.[13][14] In these organisms, the pausing induced by nucleosomes can be regulated by transcription elongation factors such as TFIIS.[14]
Elongation also involves a proofreading mechanism that can replace incorrectly incorporated bases. In eukaryotes, this may correspond with short pauses during transcription that allow appropriate RNA editing factors to bind. These pauses may be intrinsic to the RNA polymerase or due to chromatin structure.[15]

Termination[edit]
Bacteria use two different strategies for transcription termination – Rho-independent termination and Rho-dependent termination. In Rho-independent transcription termination, RNA transcription stops when the newly synthesized RNA molecule forms a G-C-rich hairpin loop followed by a run of Us. When the hairpin forms, the mechanical stress breaks the weak rU-dA bonds, now filling the DNA–RNA hybrid. This pulls the poly-U transcript out of the active site of the RNA polymerase, terminating transcription. In the 'Rho-dependent' type of termination, a protein factor called 'Rho' destabilizes the interaction between the template and the mRNA, thus releasing the newly synthesized mRNA from the elongation complex.[16]
Transcription termination in eukaryotes is less well understood than in bacteria, but involves cleavage of the new transcript followed by template-independent addition of adenines at its new 3' end, in a process called polyadenylation.[17]
Inhibitors[edit]
Transcription inhibitors can be used as antibiotics against, for example, pathogenic bacteria (antibacterials) and fungi (antifungals). An example of such an antibacterial is rifampicin, which inhibits bacterial transcription of DNA into mRNA by inhibiting DNA-dependent RNA polymerase by binding its beta-subunit, while 8-hydroxyquinoline is an antifungal transcription inhibitor.[18] The effects of histone methylation may also work to inhibit the action of transcription.
Endogenous inhibitors[edit]
In vertebrates, the majority of gene promoters contain a CpG island with numerous CpG sites.[19] When many of a gene's promoter CpG sites are methylated the gene becomes inhibited (silenced).[20] Colorectal cancers typically have 3 to 6 driver mutations and 33 to 66 hitchhiker or passenger mutations.[21] However, transcriptional inhibition (silencing) may be of more importance than mutation in causing progression to cancer. For example, in colorectal cancers about 600 to 800 genes are transcriptionally inhibited by CpG island methylation (see regulation of transcription in cancer). Transcriptional repression in cancer can also occur by other epigenetic mechanisms, such as altered expression of microRNAs.[22] In breast cancer, transcriptional repression of BRCA1 may occur more frequently by over-expressed microRNA-182 than by hypermethylation of the BRCA1 promoter (see Low expression of BRCA1 in breast and ovarian cancers).
Transcription factors[edit]
Legal Software Transcription
Active transcription units are clustered in the nucleus, in discrete sites called transcription factories or euchromatin. Such sites can be visualized by allowing engaged polymerases to extend their transcripts in tagged precursors (Br-UTP or Br-U) and immuno-labeling the tagged nascent RNA. Transcription factories can also be localized using fluorescence in situ hybridization or marked by antibodies directed against polymerases. There are ~10,000 factories in the nucleoplasm of a HeLa cell, among which are ~8,000 polymerase II factories and ~2,000 polymerase III factories. Each polymerase II factory contains ~8 polymerases. As most active transcription units are associated with only one polymerase, each factory usually contains ~8 different transcription units. These units might be associated through promoters and/or enhancers, with loops forming a 'cloud' around the factor.[23]
History[edit]
A molecule that allows the genetic material to be realized as a protein was first hypothesized by François Jacob and Jacques Monod. Severo Ochoa won a Nobel Prize in Physiology or Medicine in 1959 for developing a process for synthesizing RNA in vitro with polynucleotide phosphorylase, which was useful for cracking the genetic code. RNA synthesis by RNA polymerase was established in vitro by several laboratories by 1965; however, the RNA synthesized by these enzymes had properties that suggested the existence of an additional factor needed to terminate transcription correctly.[citation needed]
In 1972, Walter Fiers became the first person to actually prove the existence of the terminating enzyme.
Roger D. Kornberg won the 2006 Nobel Prize in Chemistry 'for his studies of the molecular basis of eukaryotic transcription'.[24]
Measuring and detecting[edit]
Transcription can be measured and detected in a variety of ways:[citation needed]
- G-Less Cassette transcription assay: measures promoter strength
- Run-off transcription assay: identifies transcription start sites (TSS)
- Nuclear run-on assay: measures the relative abundance of newly formed transcripts
- RNase protection assay and ChIP-Chip of RNAP: detect active transcription sites
- RT-PCR: measures the absolute abundance of total or nuclear RNA levels, which may however differ from transcription rates
- DNA microarrays: measures the relative abundance of the global total or nuclear RNA levels; however, these may differ from transcription rates
- In situ hybridization: detects the presence of a transcript
- MS2 tagging: by incorporating RNA stem loops, such as MS2, into a gene, these become incorporated into newly synthesized RNA. The stem loops can then be detected using a fusion of GFP and the MS2 coat protein, which has a high affinity, sequence-specific interaction with the MS2 stem loops. The recruitment of GFP to the site of transcription is visualized as a single fluorescent spot. This new approach has revealed that transcription occurs in discontinuous bursts, or pulses (see Transcriptional bursting). With the notable exception of in situ techniques, most other methods provide cell population averages, and are not capable of detecting this fundamental property of genes.[25]
- Northern blot: the traditional method, and until the advent of RNA-Seq, the most quantitative
- RNA-Seq: applies next-generation sequencing techniques to sequence whole transcriptomes, which allows the measurement of relative abundance of RNA, as well as the detection of additional variations such as fusion genes, post-transcriptional edits and novel splice sites
Reverse transcription[edit]
Some viruses (such as HIV, the cause of AIDS), have the ability to transcribe RNA into DNA. HIV has an RNA genome that is reverse transcribed into DNA. The resulting DNA can be merged with the DNA genome of the host cell. The main enzyme responsible for synthesis of DNA from an RNA template is called reverse transcriptase.
In the case of HIV, reverse transcriptase is responsible for synthesizing a complementary DNA strand (cDNA) to the viral RNA genome. The enzyme ribonuclease H then digests the RNA strand, and reverse transcriptase synthesises a complementary strand of DNA to form a double helix DNA structure ('cDNA'). The cDNA is integrated into the host cell's genome by the enzyme integrase, which causes the host cell to generate viral proteins that reassemble into new viral particles. In HIV, subsequent to this, the host cell undergoes programmed cell death, or apoptosis of T cells.[26] However, in other retroviruses, the host cell remains intact as the virus buds out of the cell.
Some eukaryotic cells contain an enzyme with reverse transcription activity called telomerase. Telomerase is a reverse transcriptase that lengthens the ends of linear chromosomes. Telomerase carries an RNA template from which it synthesizes a repeating sequence of DNA, or 'junk' DNA. This repeated sequence of DNA is called a telomere and can be thought of as a 'cap' for a chromosome. It is important because every time a linear chromosome is duplicated, it is shortened. With this 'junk' DNA or 'cap' at the ends of chromosomes, the shortening eliminates some of the non-essential, repeated sequence rather than the protein-encoding DNA sequence, that is farther away from the chromosome end.
Telomerase is often activated in cancer cells to enable cancer cells to duplicate their genomes indefinitely without losing important protein-coding DNA sequence. Activation of telomerase could be part of the process that allows cancer cells to become immortal. The immortalizing factor of cancer via telomere lengthening due to telomerase has been proven to occur in 90% of all carcinogenic tumors in vivo with the remaining 10% using an alternative telomere maintenance route called ALT or Alternative Lengthening of Telomeres.[27]
See also[edit]
- Crick's central dogma, in which the product of transcription, mRNA, is translated to form polypeptides, and where it is asserted that the reverse processes never occur
- Splicing - process of removing introns from precursor messenger RNA (pre-mRNA) to make messenger RNA (mRNA)
References[edit]
- ^ abcEldra P. Solomon, Linda R. Berg, Diana W. Martin. Biology, 8th Edition, International Student Edition. Thomson Brooks/Cole. ISBN978-0495317142
- ^Koonin EV, Gorbalenya AE, Chumakov KM (July 1989). 'Tentative identification of RNA-dependent RNA polymerases of dsRNA viruses and their relationship to positive strand RNA viral polymerases'. FEBS Letters. 252 (1–2): 42–6. doi:10.1016/0014-5793(89)80886-5. PMID2759231.
- ^'DNA Strands'. www.sci.sdsu.edu. Archived from the original on 27 October 2017. Retrieved 1 May 2018.
- ^Berg J, Tymoczko JL, Stryer L (2006). Biochemistry (6th ed.). San Francisco: W. H. Freeman. ISBN0-7167-8724-5.
- ^ abcdefgWatson JD, Baker TA, Bell SP, Gann AA, Levine M, Losick RM (2013). Molecular Biology of the Gene (7th ed.). Pearson.
- ^Roeder, Robert G. (1991). 'The complexities of eukaryotic transcription initiation: regulation of preinitiation complex assembly'. Trends in Biochemical Sciences. 16: 402–408. doi:10.1016/0968-0004(91)90164-Q. ISSN0968-0004.
- ^Goldman SR, Ebright RH, Nickels BE (May 2009). 'Direct detection of abortive RNA transcripts in vivo'. Science. 324 (5929): 927–8. doi:10.1126/science.1169237. PMC2718712. PMID19443781.
- ^Revyakin A, Liu C, Ebright RH, Strick TR (November 2006). 'Abortive initiation and productive initiation by RNA polymerase involve DNA scrunching'. Science. 314 (5802): 1139–43. doi:10.1126/science.1131398. PMC2754787. PMID17110577.
- ^Raffaelle M, Kanin EI, Vogt J, Burgess RR, Ansari AZ (November 2005). 'Holoenzyme switching and stochastic release of sigma factors from RNA polymerase in vivo'. Molecular Cell. 20 (3): 357–66. doi:10.1016/j.molcel.2005.10.011. PMID16285918.
- ^Mandal SS, Chu C, Wada T, Handa H, Shatkin AJ, Reinberg D (May 2004). 'Functional interactions of RNA-capping enzyme with factors that positively and negatively regulate promoter escape by RNA polymerase II'. Proceedings of the National Academy of Sciences of the United States of America. 101 (20): 7572–7. doi:10.1073/pnas.0401493101. PMC419647. PMID15136722.
- ^Goodrich JA, Tjian R (April 1994). 'Transcription factors IIE and IIH and ATP hydrolysis direct promoter clearance by RNA polymerase II'. Cell. 77 (1): 145–56. doi:10.1016/0092-8674(94)90242-9. PMID8156590.
- ^Milo, Ron; Philips, Rob. 'Cell Biology by the Numbers: What is faster, transcription or translation?'. book.bionumbers.org. Archived from the original on 20 April 2017. Retrieved 8 March 2017.
- ^Hodges C, Bintu L, Lubkowska L, Kashlev M, Bustamante C (July 2009). 'Nucleosomal fluctuations govern the transcription dynamics of RNA polymerase II'. Science. 325 (5940): 626–8. doi:10.1126/science.1172926. PMC2775800. PMID19644123.
- ^ abFitz V, Shin J, Ehrlich C, Farnung L, Cramer P, Zaburdaev V, Grill SW (2016). 'Nucleosomal arrangement affects single-molecule transcription dynamics'. Proceedings of the National Academy of Sciences. 113 (45): 12733–12738. PMC5111697. Archived from the original on 2018-05-01.
- ^Yukihara; et al. (1985). 'Eukaryotic transcription: a summary of research and experimental techniques'. Journal of Molecular Biology. 14 (21): 56–79.
- ^Richardson JP (September 2002). 'Rho-dependent termination and ATPases in transcript termination'. Biochimica et Biophysica Acta. 1577 (2): 251–260. doi:10.1016/S0167-4781(02)00456-6. PMID12213656.
- ^Lykke-Andersen S, Jensen TH (October 2007). 'Overlapping pathways dictate termination of RNA polymerase II transcription'. Biochimie. 89 (10): 1177–82. doi:10.1016/j.biochi.2007.05.007. PMID17629387.
- ^8-Hydroxyquinoline info from SIGMA-ALDRICH. Retrieved Feb 2012
- ^Saxonov S, Berg P, Brutlag DL (January 2006). 'A genome-wide analysis of CpG dinucleotides in the human genome distinguishes two distinct classes of promoters'. Proceedings of the National Academy of Sciences of the United States of America. 103 (5): 1412–7. doi:10.1073/pnas.0510310103. PMC1345710. PMID16432200.
- ^Bird A (January 2002). 'DNA methylation patterns and epigenetic memory'. Genes & Development. 16 (1): 6–21. doi:10.1101/gad.947102. PMID11782440.
- ^Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA, Kinzler KW (March 2013). 'Cancer genome landscapes'. Science. 339 (6127): 1546–58. doi:10.1126/science.1235122. PMC3749880. PMID23539594.
- ^Tessitore A, Cicciarelli G, Del Vecchio F, Gaggiano A, Verzella D, Fischietti M, Vecchiotti D, Capece D, Zazzeroni F, Alesse E (2014). 'MicroRNAs in the DNA Damage/Repair Network and Cancer'. International Journal of Genomics. 2014: 820248. doi:10.1155/2014/820248. PMC3926391. PMID24616890.
- ^Papantonis A, Kohro T, Baboo S, Larkin JD, Deng B, Short P, Tsutsumi S, Taylor S, Kanki Y, Kobayashi M, Li G, Poh HM, Ruan X, Aburatani H, Ruan Y, Kodama T, Wada Y, Cook PR (November 2012). 'TNFα signals through specialized factories where responsive coding and miRNA genes are transcribed'. The EMBO Journal. 31 (23): 4404–14. CiteSeerX10.1.1.919.1919. doi:10.1038/emboj.2012.288. PMC3512387. PMID23103767.
- ^'Chemistry 2006'. Nobel Foundation. Archived from the original on March 15, 2007. Retrieved March 29, 2007.
- ^Raj A, van Oudenaarden A (October 2008). 'Nature, nurture, or chance: stochastic gene expression and its consequences'. Cell. 135 (2): 216–26. doi:10.1016/j.cell.2008.09.050. PMC3118044. PMID18957198.
- ^Kolesnikova IN (2000). 'Some patterns of apoptosis mechanism during HIV-infection'. Dissertation (in Russian). Archived from the original on July 10, 2011. Retrieved February 20, 2011.
- ^Cesare AJ, Reddel RR (May 2010). 'Alternative lengthening of telomeres: models, mechanisms and implications'. Nature Reviews. Genetics. 11 (5): 319–30. doi:10.1038/nrg2763. PMID20351727.
External links[edit]
| Wikimedia Commons has media related to Transcription (genetics). |
Alibaba Software Transcription Factory
- Interactive Java simulation of transcription initiation. From Center for Models of Life at the Niels Bohr Institute.
- Interactive Java simulation of transcription interference--a game of promoter dominance in bacterial virus. From Center for Models of Life at the Niels Bohr Institute.